From Cat Photos to Colonoscopies
Virgo — Leaders in GI Endoscopy Data Launched in 2018, Virgo is now the leading cloud video capture and management platform for endoscopy.
Virgo — Leaders in GI Endoscopy Data
Launched in 2018, Virgo is now the leading cloud video capture and management platform for endoscopy. Today, we’re proud to support physicians in the US and abroad — including many of US News and World Reports’ Top 20 gastroenterology programs. In doing so, Virgo has amassed the world’s largest dataset of endoscopy videos, which continues to grow at an ever-increasing rate.
Early Bet on Data Infrastructure
But our success story isn't just about scale; it's about strategic foresight. Today’s AI hype isn’t unfamiliar to Virgo. When we were raising our first round of capital in 2017, there was plenty of AI buzz in healthcare, particularly in gastroenterology. Instead of jumping on the AI bandwagon, we made an early bet to focus on data infrastructure. We recognized that AI capabilities would continue democratizing faster than any individual startup was likely to advance. Instead, we focused on building the data infrastructure sorely missing in endoscopy. Before Virgo, it was exceptionally rare for doctors to save procedure videos. A handful of still images would be committed to the medical record, but from a big data and AI perspective, this lost data represents a massive opportunity. Data infrastructure is key because it both provides the most necessary resource to build AI systems and also affords a place to then deploy these AI systems. With the recent rapid advances in foundation models for computer vision, our bet is finally paying off.
AI Foundation Models and Endoscopy
Foundation models are AI models trained on a large amount of data, often in a self-supervised manner. Once trained, foundation models can then be adapted to help solve specific tasks. This process generally reduces the computation and annotation needs for new task development that would otherwise be required with traditional supervised learning. Some of the most well-known examples of foundation models are large language models (LLMs) like GPT-3 and GPT-4 (which power ChatGPT) and computer vision models like CLIP (which powers Stable Diffusion). In a recent development that underscores the democratization of AI technology, Meta has open-sourced a new computer vision foundation model called DINOv2. This computer vision model has been trained using self-supervision, a method that doesn't require manually labeled data. The stats are remarkable — DINOv2 was trained using 142 million unlabelled images. This makes it incredibly versatile and adaptable to new use cases. Out of the box, DINOv2 can rapidly jumpstart all sorts of computer vision tasks, including image classification, object detection, and depth estimation. And this has massive implications for AI development in healthcare.
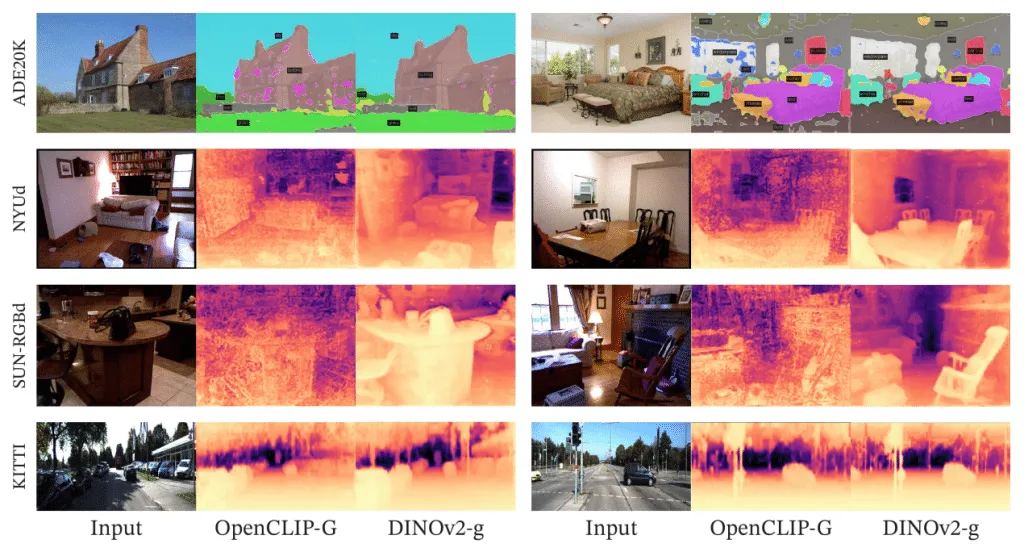

Segmentation and depth estimation with linear classifiers. Examples from ADE20K, NYUd, SUN RGB-D and KITTI with a linear probe on frozen OpenCLIP-G and DINOv2-g features. Source: DINOv2: Learning Robust Visual Features without Supervision
Over the past few weeks, we’ve been evaluating using DINOv2 for endoscopy. While it is unlikely that the DINOv2 training data contained significant quantities of endoscopy images, we are already seeing excellent results in endoscopy. One of the problems we face with our dataset is the scale and growth of unlabelled data. Every day we ingest thousands of new full-length endoscopy videos. Given the size of our team, we simply don’t have the resources to label every frame or even every video manually. DINOv2 to the rescue. We’ve found that out-of-the-box, the DINOv2 feature embeddings are able to powerfully cluster concepts within endoscopy. Using a t-distributed stochastic neighbor embedding (t-SNE) plot, we quickly visualized how the DINOv2 features represented the images. Here is a plot of sampled frames from a handful of different colonoscopy videos:
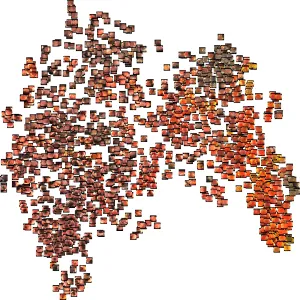
Even at first glance, you get the sense that the DINOv2 features represent…something. It appears as though frames are perhaps grouped roughly by color. Zooming in, though, we see that deeper representations also exist. Toward the lower right portion of the image, we see a cluster of frames that are fairly homogeneous in color, with little actual definition. This typically occurs when the tip of the endoscope sits flush up against the colon wall.
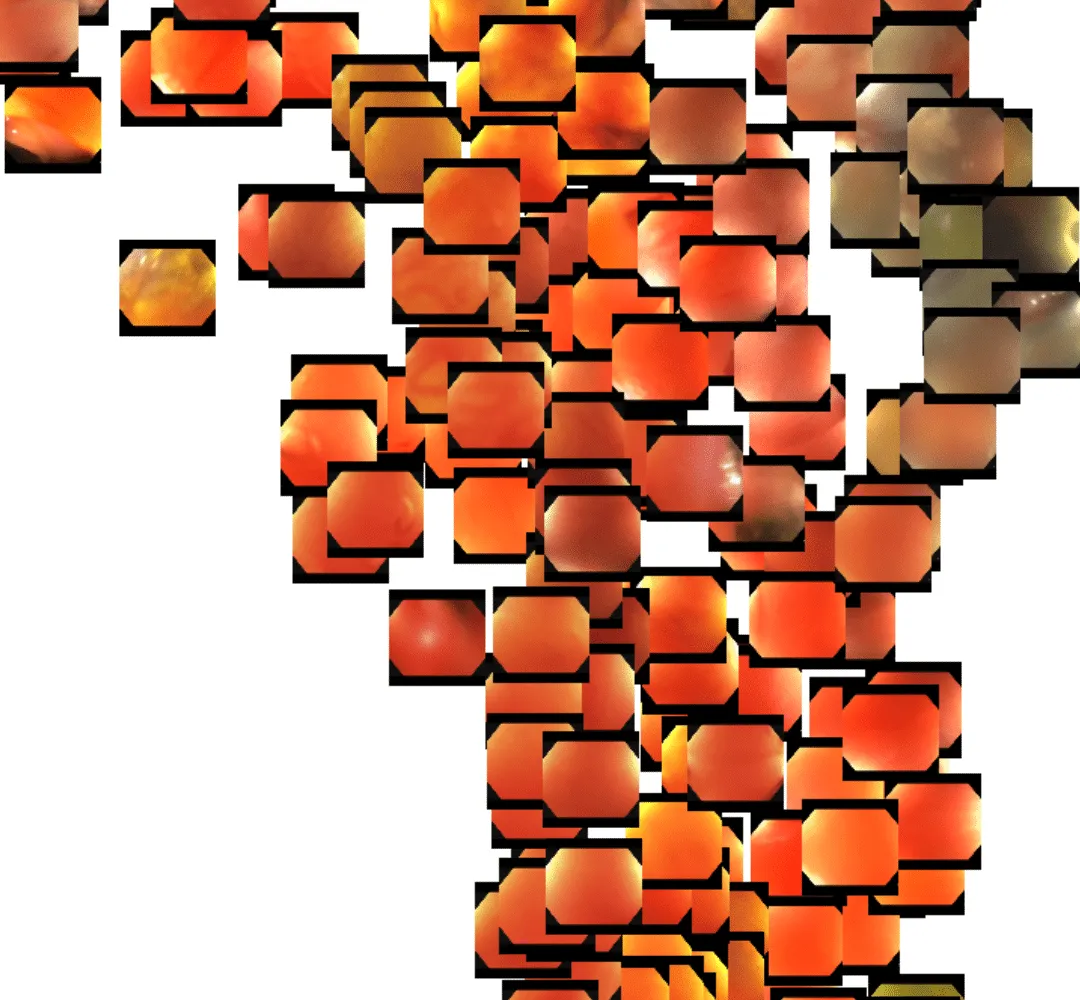

In the upper right, we see a cluster of similar images but with a bit more reflective texture. This is usually indicative of the endoscopist flushing the colon with rushing water.
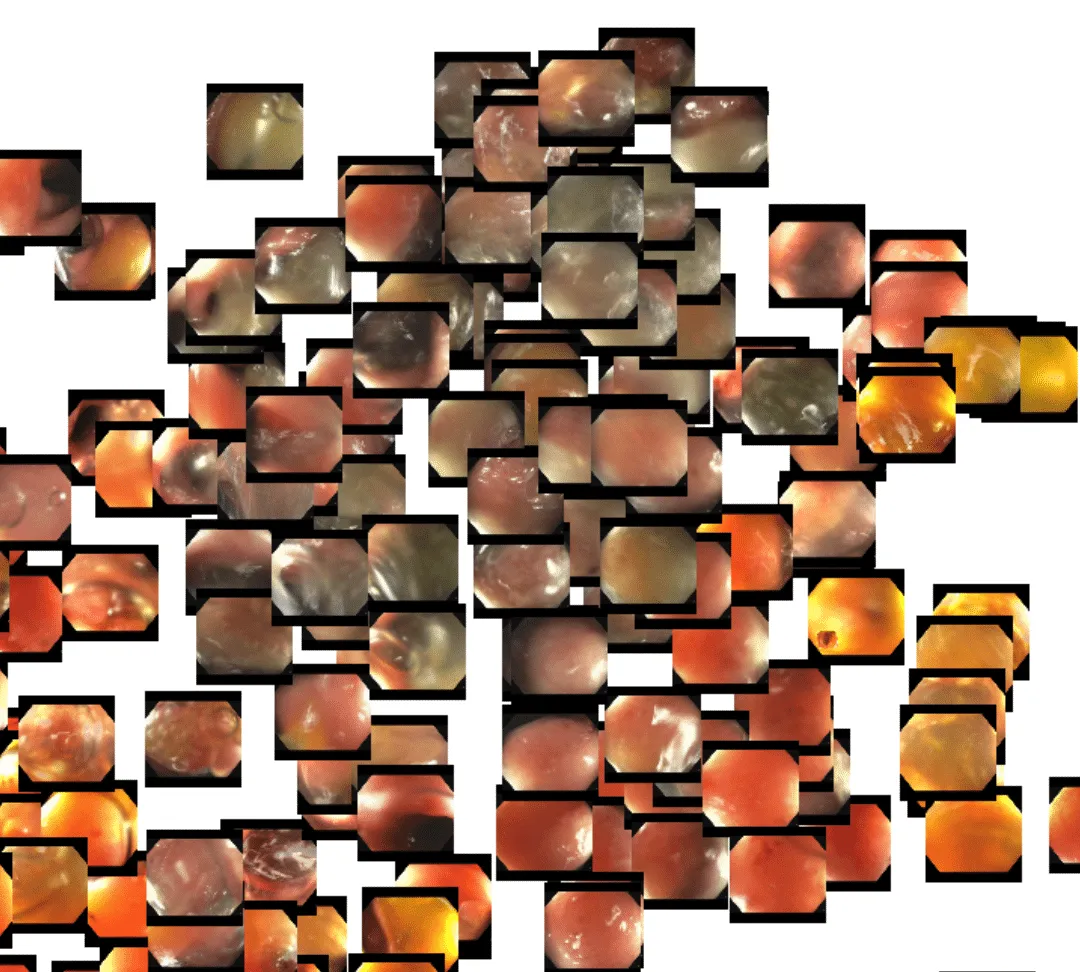

Here in the upper left, we see a cluster of related frames that have biopsy forceps present in view:
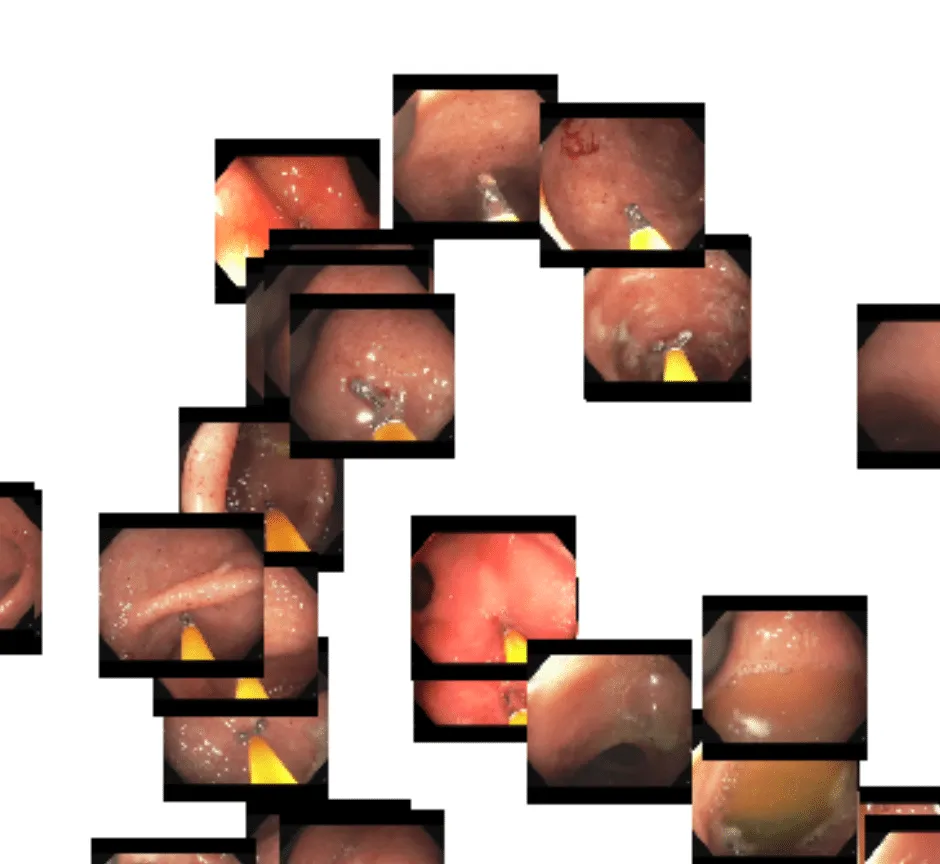
And here is a cluster of frames containing a snare being used to resect a potentially cancerous polyp:
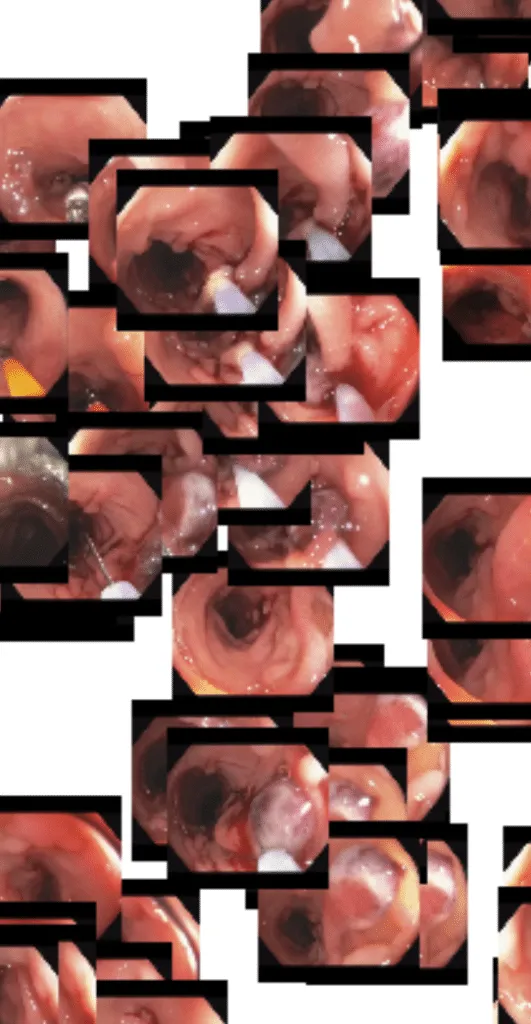
Finally, we are particularly interested in a cluster toward the bottom of the image — frames containing inflammation and ulceration. These are potentially indicative of inflammatory bowel disease and may be useful for identifying possible candidates for clinical trials.
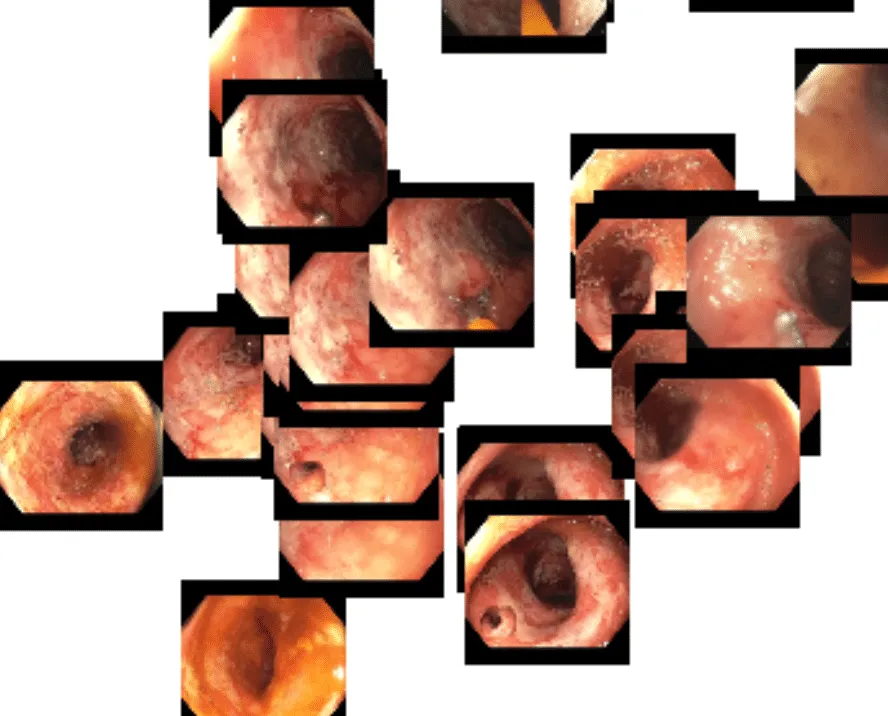
While this is an interesting exercise in its own right, the most compelling aspect is that all of this clustering is being achieved using a machine learning model that was trained on 142 million non-endoscopic images. This was our first indication that DINOv2 could be a powerful foundation model for our AI development in endoscopy.
From Foundation Model to Instrument Detection
Virgo’s database of endoscopy procedure videos is ideally suited to take advantage of this foundation model paradigm. Every day, we ingest massive amounts of unlabeled endoscopy video data and add it to our even more massive trove of unlabeled endoscopy videos. In a fully supervised learning paradigm, we would need to hire or pay experts to manually review these videos and then painstakingly label any individual frames. This can be expensive and time-consuming, often to the point of being resource-prohibitive for startups. One task where we’ve already explored comparing the foundation model and supervised learning paradigms is instrument detection. During a colonoscopy (or other endoscopic procedure), doctors aren’t just looking for something abnormal to make a diagnosis. Often, they’re also looking to actually do something — biopsy the mucosal wall to send to pathology, resect a polyp with a snare, tattoo a tumor for identification by a surgeon, or dilate a stricture in the colon with a balloon.
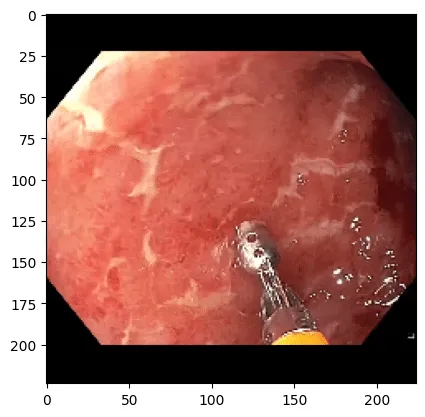
Example of forceps being used to biopsy what appears to be inflamed and ulcerated colon mucosa.
In all of these situations, an instrument is placed through a long channel in the endoscope so that it enters the field of view and can be operated by the endoscopist. For Virgo, instrument detection is a useful feature because we can overlay information about a video that enables a physician to quickly hone in on sections of a longer video that are most meaningful without them having to watch the entire thing. Currently, Virgo offers detection of snares and forceps, which helps doctors find sections of the video where biopsies and polypectomies occurred.
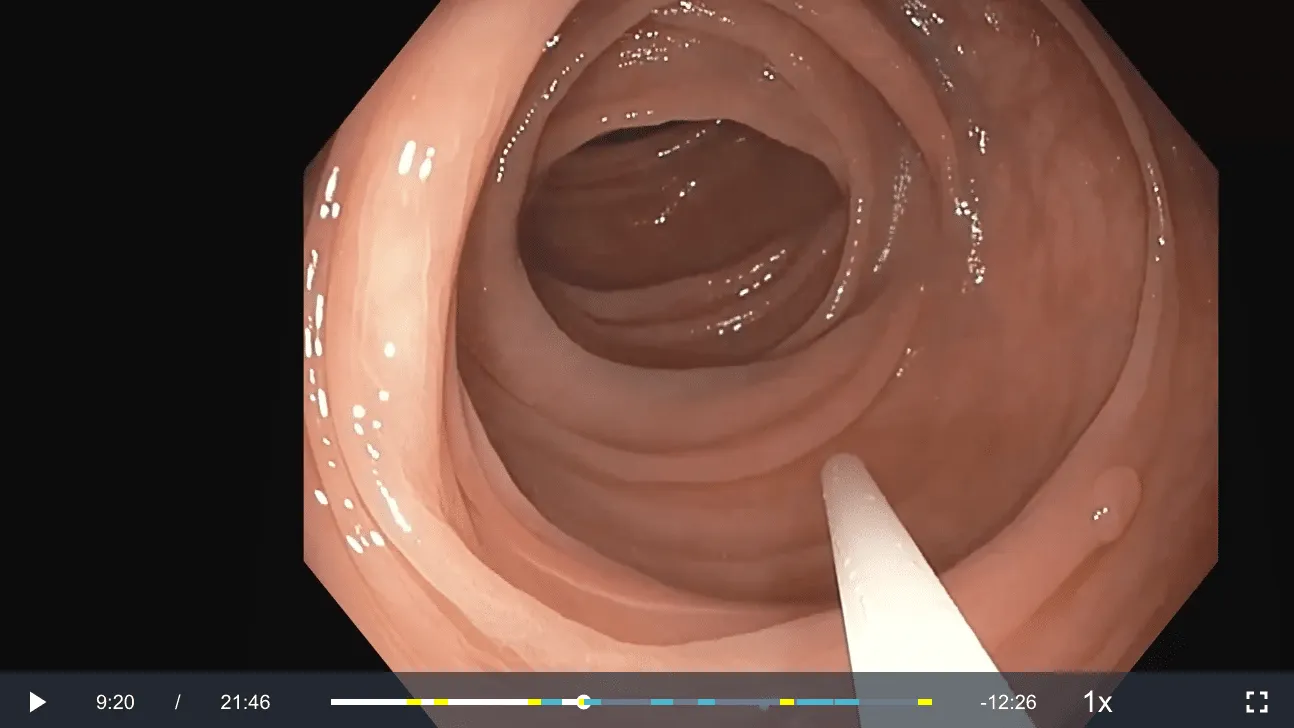
Example of a video in the Virgo platform where a snare was identified and represented on the video timeline by a blue marker. Here the snare is being inserted to resect the polyp in the lower right portion of the image.
Virgo’s current instrument detection system was trained in a fully supervised paradigm. We labeled tens of thousands of individual frames from 500 different colonoscopy videos. Then, we trained a convolutional neural network model to classify frames with either no instrument, a snare, or a forceps. The results are quite good and result in a very useful product, but this took us several weeks of internal labeling, experimentation, and development. On top of that, we would love to expand our instrument detection capabilities in several ways — more instrument types, better accuracy, etc.; however, with limited resources and a small team, the prospect of labeling 10’s of thousands of additional frames is daunting. Enter DINOv2 as a foundation model. We’ve found that the extracted features from the DINOv2 model are so good that we can train new models for essentially any task we can conceive of with significantly less labeled data. This works particularly well in an active learning environment, where we regularly retrain models and use the newest model to help focus attention on the most important data to label.
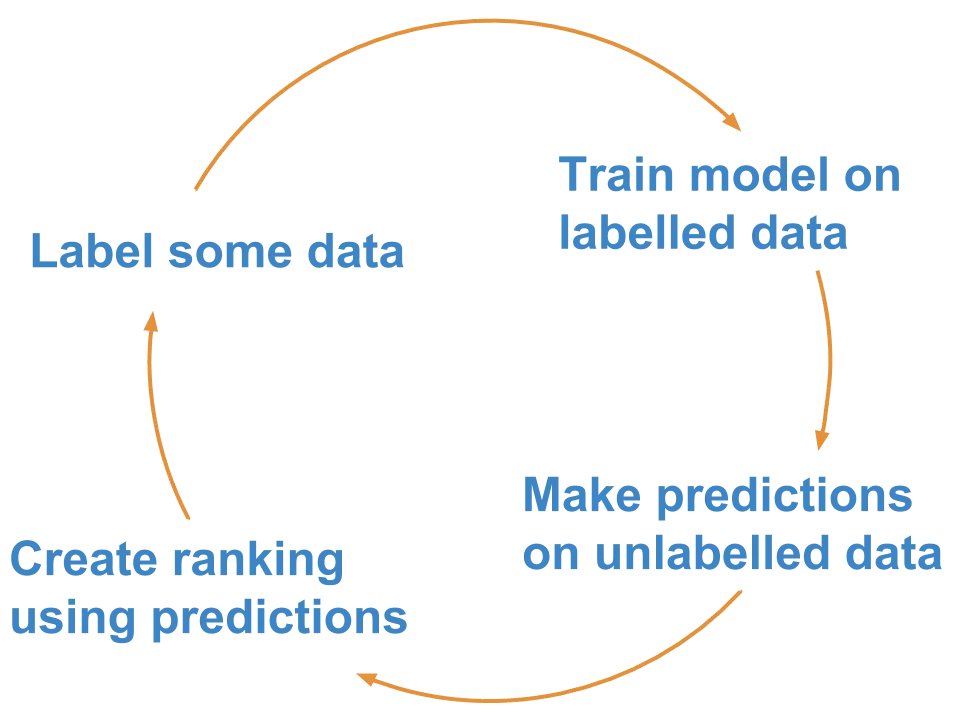
Rough steps of an active learning framework. Source: Superintendent Python library
To give an example of how this works, we’ll look again at instrument detection — this time expanded to include the following classes:
- None
- Retroflexion (when the scope looks back at itself)
- Snare (used to resect polyps)
- Forcep (used to resect small polyps and biopsy tissue)
- Net (used to retrieve resected polyps)
- Needle (used to inject tattoo dye and lifting agent)
- Clip (used to stop bleeding)
- Knife (used to cut tissue with electrocautery)
- Guidewire (used to cannulate narrow spaces)
- Sphincterotome (used to make small cuts in strictures)
- Balloon (used to dilate strictures)
- Stent (used to create a permanent portal from one area of the body to another)
In this framework, we simply sample a random set of videos from the broader Virgo dataset, extract frames from these videos, and then extract the DINOv2 features from these frames. We then trained a simple linear classifier using these features. As a proof of concept, we set up an active learning annotation environment as shown here using the ipyannotations and superintendent libraries:
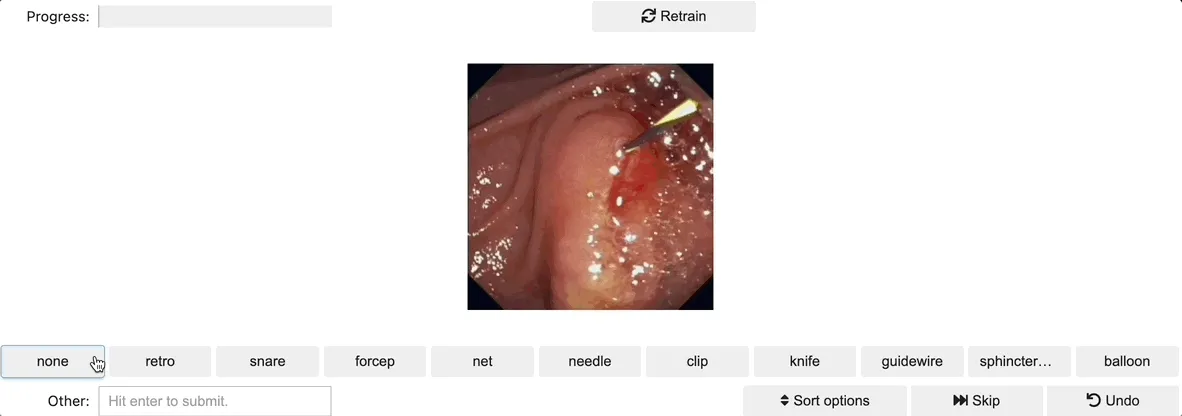
We conducted the first training run after labeling just a handful of images. Superintendent then automatically handles serving up images based on your trained model and selected acquisition function. In doing so, you spend time labeling data that theoretically will be most informative for refining your model. In practice, we’ve found that for multiple tasks, this methodology with the DINOv2 features leads very quickly to models that perform at least well enough to direct active learning productively.
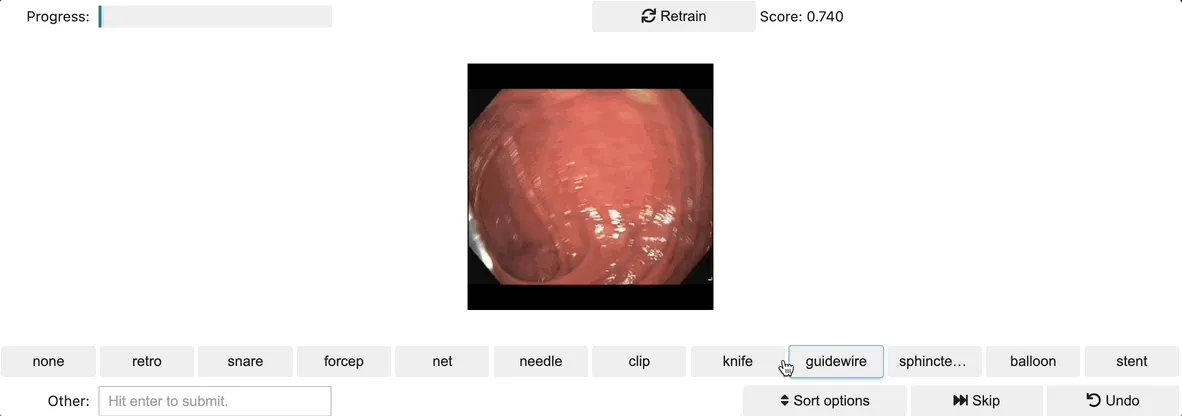
With only a few dozen images labeled, the model started performing decently well, and superintendent was pretty clearly serving up images that would be meaningful for refining the model further. After labeling just a few hundred images, we already had a model that felt just as, if not more, performant than our current production instrument detection; plus, this includes nine additional instrument types. Proof of concept, complete!
Superpowers for Small Teams
DINOv2 was released and open-sourced on April 17, 2023. In the intervening weeks, it has proven to be a game-changer for the team at Virgo. Over the past few weeks, we have successfully demonstrated proofs of concept for all sorts of workflow and diagnostic tasks. Our next set of challenges includes experimenting on how model size impacts performance and extending DINOv2 embeddings across video time sequences. In time though, we believe this architecture will underscore all AI development at Virgo. The really fun part is that while the team at Virgo is relatively small, we are also extremely capable, and tools like DINOv2 give us new superpowers. Startups like Virgo are nimble and perfectly positioned to take advantage of ongoing advancements in foundation models. If you’re an engineer, researcher, physician, or pharma company who is interested in working with our dynamic team, please email me at matt@virgosvs.com!
Virgo's Vision
So what’s next? We're excited about the potential to use DINOv2 to improve diagnostic accuracy, identify rare conditions, accelerate clinical trials, support pharmaceutical R&D, and help doctors provide better care. But this is just the beginning. We are also increasingly optimistic about foundation models in general and their development for specific domains. A particularly interesting section of the DINOv2 paper poses the question of whether or not the model needs to be fine-tuned. In their experimentation, the Meta team found that fine-tuning provided marginal benefit, but in general it’s optional.
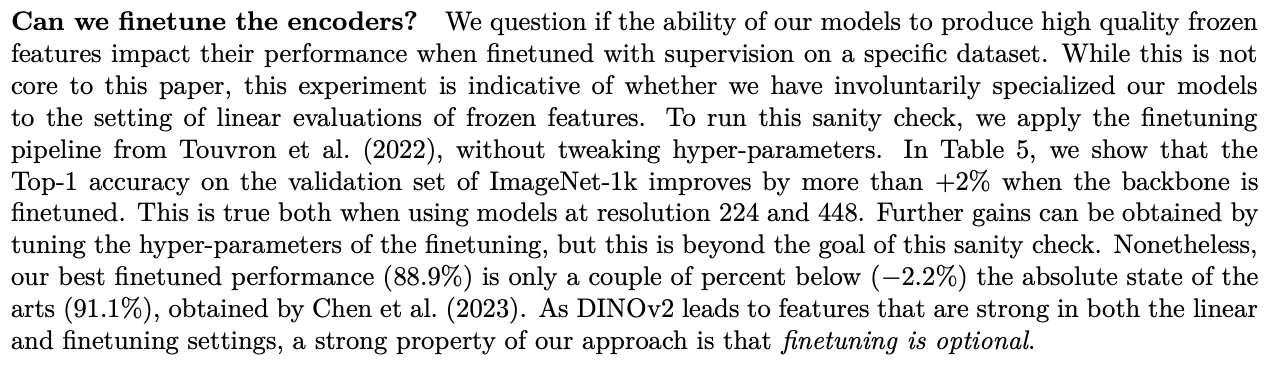

That said, this evidence of marginal benefit is quite intriguing, particularly as it relates to niche applications such as endoscopy. We think there’s a good chance that a new crop of verticalized foundation models will emerge and drive significant progress in their respective fields. Virgo is actively exploring development of an endoscopy-specific foundation model on top of our growing dataset. This is likely an area of significant opportunity for other startups that are sitting on unique data, infrastructure, and insights. There are also very exciting opportunities emerging for startups to take advantage of GPU clusters for training. Two that are particularly worth calling out are from Lambda Labs and the Nat Friedman / Daniel Gross duo.
We fully expect an explosion of startups taking advantage of these types of resources to develop new open and closed foundation models. We look forward to being a part of that future! Again, if you’re an engineer or researcher interested in collaborating on the development of endoscopy foundation models, or if you’re a GI provider or pharmaceutical company and at all interested in what we’re building here at Virgo, we would love to hear from you. Find us on Twitter, LinkedIn, or email me directly at matt@virgosvs.com.
Acknowledgments
Thanks to Nikhil Krishnan, Alex Iskold, Timothée Darcet, Sanjana Basu, Shubhra Jain, Eric Norlin, and Rob May for providing feedback during the writing of this post.
Share this post